If there is a video gamer in your life, chances are that you have heard of Spore, the latest creation from the super successful inventor of ‘The Sims’.
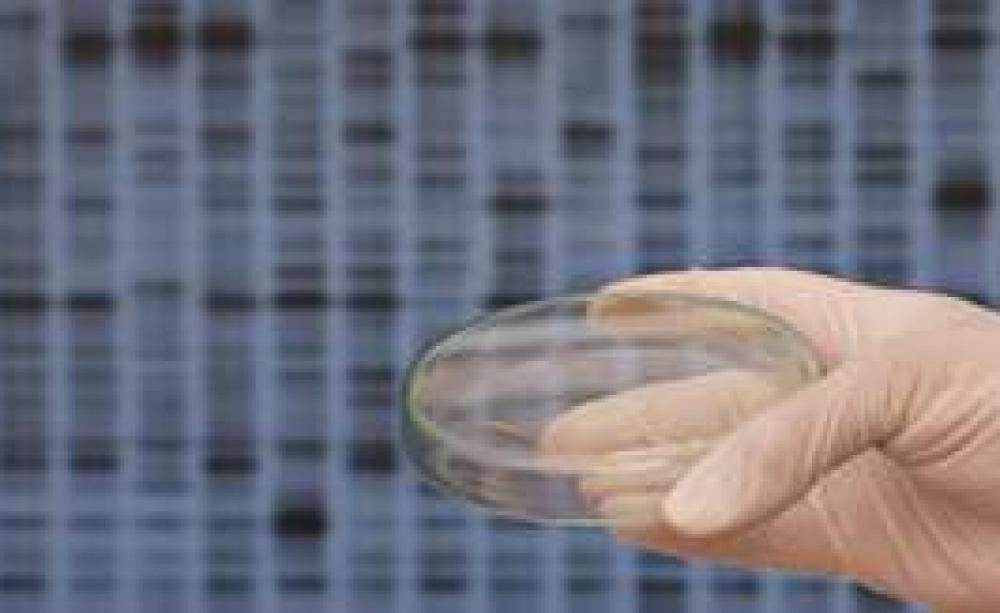
Spore lets players digitally design and evolve new organisms ranging from single-celled microbes to intergalactic aliens. In the game, the user can genetically alter their digital life-form, adding body parts one click at a time. The user-created organisms are simultaneously released into the game-worlds of thousands of other players, creating complex and surprising ecosystems to navigate. Spore organisms can be printed on to a t-shirt, mailed to you as plastic fi gurines or uploaded to ‘Sporepedia’, an online gallery where fans share their custom-made life-forms for remixing. Bored with editing digital music and digital video, the next nerd frontier is digital life.
As would-be intelligent designers experiment with Spore, the line between remixing digital and biological life is becoming perilously thin. Thanks to the new tools of synthetic biology, Spore fans wanting to dabble in a more organic medium can already do so from the comfort of their laptops. At www.biobricks.org, you can choose among thousands of custom-designed genetic sequences (known as ‘standard biological parts’) that can be subsequently posted to you as real DNA. One startup, Ginkgo BioWorks of Boston, will soon be offering kits of DNA stuck to paper that enable simple genetic engineering at home. Drop the paper in a test tube with some nutrients and bacteria. Chill it in the fridge and then warm it against your computer screen. Presto! Your bacteria are genetically altered to glow red or smell like bananas.
No lab bench or white coat in sight. Just as video production moved from a niche profession requiring expensive equipment to today’s universe of amateur YouTube clips, so a new generation of synthetic biologists is working to take the elite science of genetic engineering away from Monsanto and give it to the masses as a craft.
They call this DIY genetic engineering ‘garage biology’ or ‘biohacking’. In Boston and San Francisco, amateur biohacking clubs are beginning to form whose aim is to replicate the success of the computer industry in kick-starting an industrial revolution by bringing technology into the home. Synthetic biologist Drew Endy of Stanford University imagines that within a few years there will be a whole new class of life-form designers working remotely from their laptops – emailing their designs to labs just as graphic designers send digital files for printing. Endy runs an annual competition in which hundreds of students and teenagers compete to create the ‘coolest’ life-form out of standard parts. Right now they ‘hack’ bacteria and yeast to take photos or secrete biofuels. Within five years, biohacking plants and animals this way will be more common.
Futurist Freeman Dyson argues that the coming outpouring of new synthetic life-forms designed by hobbyist amateurs may eventually outnumber those species developed through natural evolution, with engineered life rapidly becoming the norm.
Dyson may be overstating things, but there is certainly a big shift under way that even activists and regulators have yet to comprehend. Despite millions of acres of genetically modified corn and soya, the actual number of new species developed through genetic engineering has so far been a tiny trickle compared to the flood of engineered species our ecosystems may be about to experience. Existing biosafety regulations are already barely able to monitor the health and environmental impacts of that trickle, let alone the results of amateur biohacking carried out in kitchens, bathrooms, garages and via the internet. Unlike computing, the haphazard results of this biological programming will be living, self-replicating organisms.
Unfortunately, the digital world doesn’t seem to offer much in the way of control solutions. If the DIY organisms of Spore should go awry there is always the option of pulling the plug and rebooting the computer. Out here in the real world, that option doesn’t look so attractive.
This article first appeared in the Ecologist October 2008